fBm-Based Generative Inpainting for the Reconstruction of Chromosomal Distances
Authors: Alexander Lobashev, Dmitry Guskov, Kirill Polovnikov
🎯 Key Contributions
- Applies diffusion-based inpainting to reconstruct chromosomal distance matrices
- Demonstrates effective reproduction of fBm correlations across different memory regimes
- Outperforms standard bioinformatics approaches for FISH data imputation
Abstract
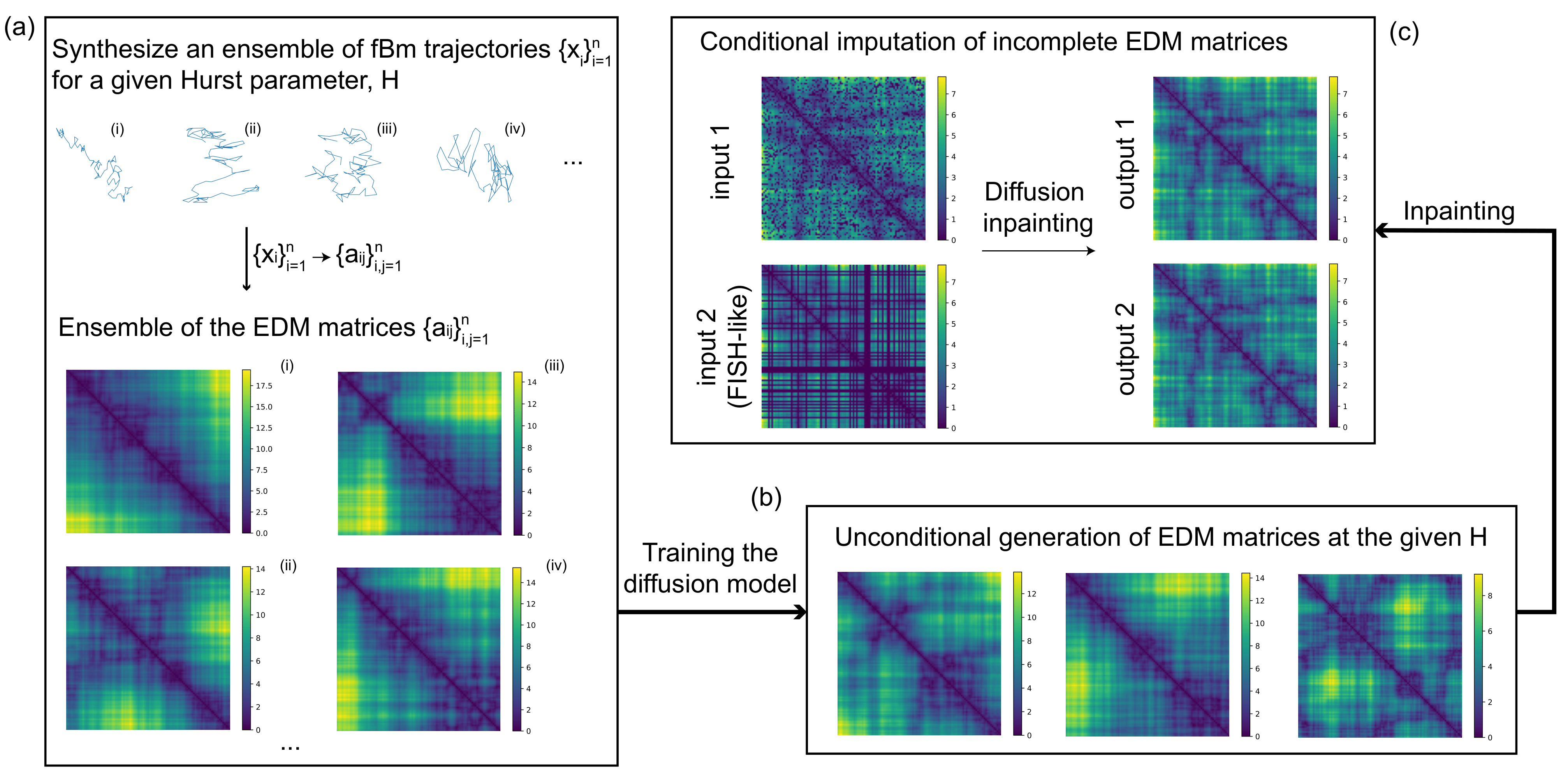
Fractional Brownian motion (fBm) features both randomness and strong scale-free correlations, challenging generative models to reproduce the intrinsic memory characterizing the underlying stochastic process.
Here we examine a zoo of diffusion-based inpainting methods on a specific dataset of corrupted images, which represent incomplete Euclidean distance matrices (EDMs) of fBm at various memory exponents. Our dataset implies uniqueness of the data imputation in the regime of low missing ratio, providing the unique ground truth for the inpainting.
We find that conditional diffusion generation readily reproduces the built-in correlations of fBm paths in different memory regimes (i.e., for sub-, Brownian, and super-diffusion trajectories), providing a robust tool for statistical imputation at high missing ratios.
As a biological application, we apply our fBm-trained diffusion model for the imputation of microscopy-derived distance matrices of chromosomal segments (Fluorescence In Situ Hybridization data)—incomplete due to experimental imperfections—and demonstrate its superiority over the standard approaches used in bioinformatics.
📋 Citation
@inproceedings{lobashev2024fbm,
title={fBm-Based Generative Inpainting for the Reconstruction of Chromosomal Distances},
author={Lobashev, Alexander and Guskov, Dmitry and Polovnikov, Kirill},
booktitle={Machine Learning for Physical Sciences Workshop at NeurIPS},
year={2024}
}